Introduction to SSM Analysis¶
This notebook provides an introduction to Structural Summary Method (SSM) analysis using the circumplex package.
1. Loading Example Data: jz2017¶
We’ll use the jz2017 dataset from Zimmermann & Wright (2017). This
dataset includes self-report data from 1166 undergraduate students who
completed:
- A circumplex measure of interpersonal problems (IIP-SC) with eight subscales: PA, BC, DE, FG, HI, JK, LM, NO
- A measure of personality disorder symptoms with ten subscales: PARPD, SCZPD, SZTPD, ASPD, BORPD, HISPD, NARPD, AVPD, DPNPD, OCPD
| Gender | PA | BC | DE | FG | HI | JK | LM | NO | PARPD | SCZPD | SZTPD | ASPD | BORPD | HISPD | NARPD | AVPD | DPNPD | OCPD |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Female | 1.5 | 1.5 | 1.25 | 1.0 | 2.0 | 2.5 | 2.25 | 2.5 | 4 | 3 | 7 | 7 | 8 | 4 | 6 | 3 | 4 | 6 |
| Female | 0.0 | 0.25 | 0.0 | 0.25 | 1.25 | 1.75 | 2.25 | 2.25 | 1 | 0 | 2 | 0 | 1 | 2 | 3 | 0 | 1 | 0 |
| Female | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 | 1 | 0 | 4 | 1 | 5 | 4 | 0 | 0 | 1 |
| Male | 2.0 | 1.75 | 1.75 | 2.5 | 2.0 | 1.75 | 2.0 | 2.5 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
| Female | 0.25 | 0.5 | 0.25 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 |
2. Mean-Based SSM Analysis¶
Single Group Analysis¶
Let’s analyze the interpersonal problems of the average individual in
the dataset using ssm_analyze(). The function returns an SSM object
with results, scores, and analysis details.
# Define circumplex scales and their angular positions
scales = ["PA", "BC", "DE", "FG", "HI", "JK", "LM", "NO"]
angles = [90, 135, 180, 225, 270, 315, 360, 45]
# Run SSM analysis
results = ssm_analyze(data=jz2017, scales=scales, angles=angles)
# Display summary
results.summary()
Statistical Basis: Mean Scores Bootstrap Resamples: 2000 Confidence Level: 0.95 Listwise Deletion: True Scale Displacements: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0] Profile[All] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.917 │ 0.889 │ 0.945 │ │ X-Value │ 0.351 │ 0.323 │ 0.378 │ │ Y-Value │ -0.252 │ -0.281 │ -0.223 │ │ Amplitude │ 0.432 │ 0.402 │ 0.46 │ │ Displacement │ 324.292 │ 320.757 │ 327.79 │ │ Model Fit │ 0.878 │ │ │ └──────────────┴──────────┴──────────┴──────────┘
Accessing Results¶
The SSM object provides easy access to results, scores, and analysis details:
# View results DataFrame with parameter estimates and confidence intervals
GT(results.results.round(2))
| Label | Group | Measure | e_est | e_lci | e_uci | x_est | x_lci | x_uci | y_est | y_lci | y_uci | a_est | a_lci | a_uci | d_est | d_lci | d_uci | fit_est |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| All | All | 0.92 | 0.89 | 0.95 | 0.35 | 0.32 | 0.38 | -0.25 | -0.28 | -0.22 | 0.43 | 0.4 | 0.46 | 324.29 | 320.76 | 327.79 | 0.88 |
| Label | PA | BC | DE | FG | HI | JK | LM | NO |
|---|---|---|---|---|---|---|---|---|
| All | 0.55 | 0.58 | 0.62 | 0.76 | 1.21 | 1.21 | 1.48 | 0.9 |
# View analysis details
print(f"Analysis type: {results.type}")
print(f"Angles: {results.details.angles}")
print(f"Bootstrap resamples: {results.details.boots}")
print(f"Confidence level: {results.details.interval}")
Analysis type: mean
Angles: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0]
Bootstrap resamples: 2000
Confidence level: 0.95
3. Visualizing SSM Results¶
The circumplex package provides three visualization functions:
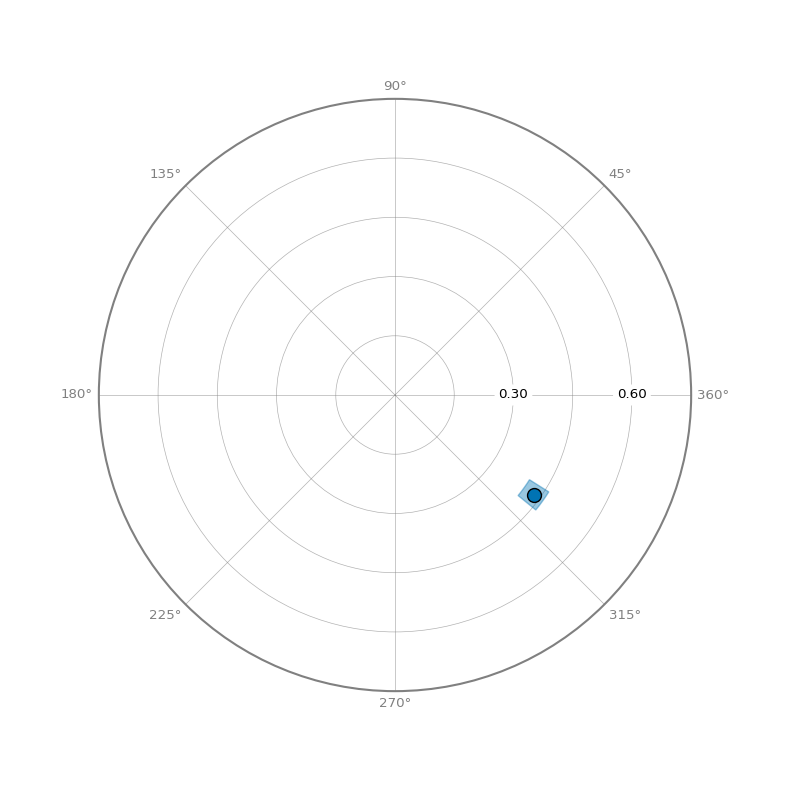
- Circle Plot: Shows amplitude and displacement on a circular plot
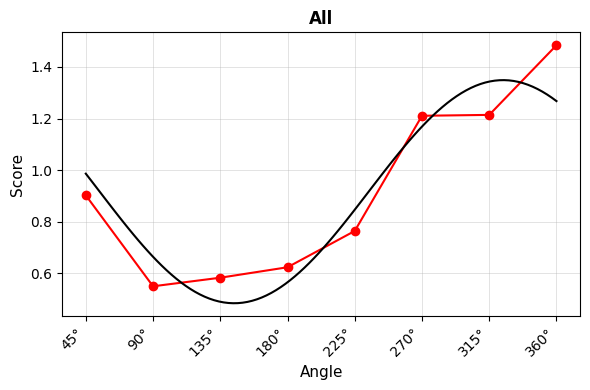
- Curve Plot: Shows fitted cosine curves overlaid on observed scores
- Contrast Plot: Shows parameter differences between groups (for contrast analyses)
Circle Plot¶
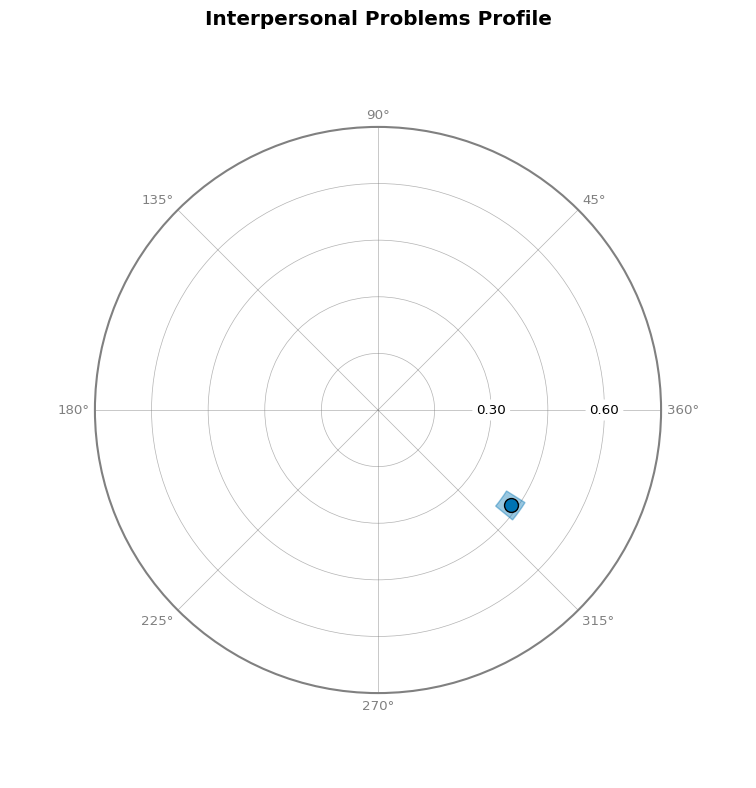
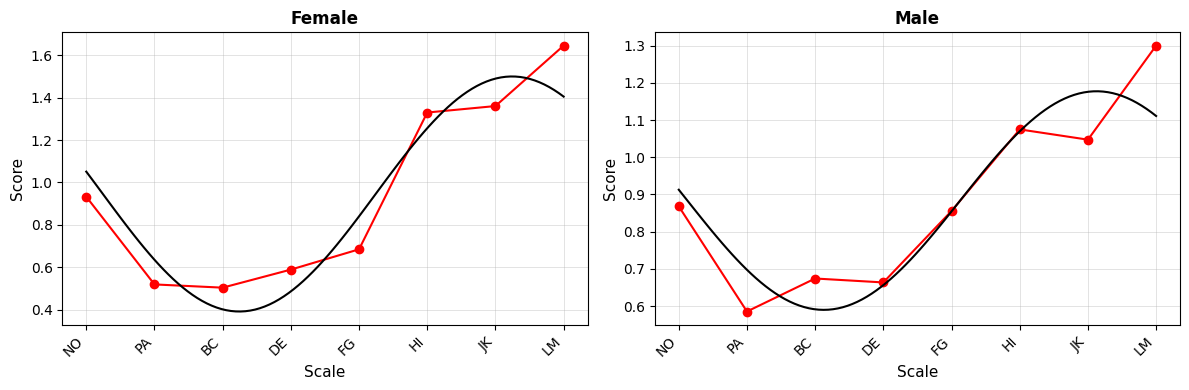
Curve Plot¶
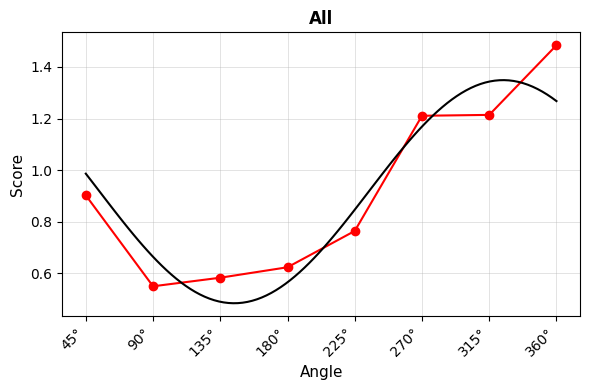
4. Correlation-Based SSM Analysis¶
To examine how personality disorder symptoms relate to interpersonal problems, we can perform correlation-based SSM analysis by specifying external measures.
# Analyze narcissistic personality disorder symptoms
results_corr = ssm_analyze(
data=jz2017,
scales=scales,
angles=angles,
measures="NARPD",
)
results_corr.summary()
Statistical Basis: Correlation Scores Bootstrap Resamples: 2000 Confidence Level: 0.95 Listwise Deletion: True Scale Displacements: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0] Profile[NARPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.202 │ 0.167 │ 0.237 │ │ X-Value │ -0.062 │ -0.094 │ -0.029 │ │ Y-Value │ 0.179 │ 0.145 │ 0.212 │ │ Amplitude │ 0.189 │ 0.155 │ 0.224 │ │ Displacement │ 108.967 │ 99.221 │ 118.54 │ │ Model Fit │ 0.957 │ │ │ └──────────────┴──────────┴──────────┴──────────┘
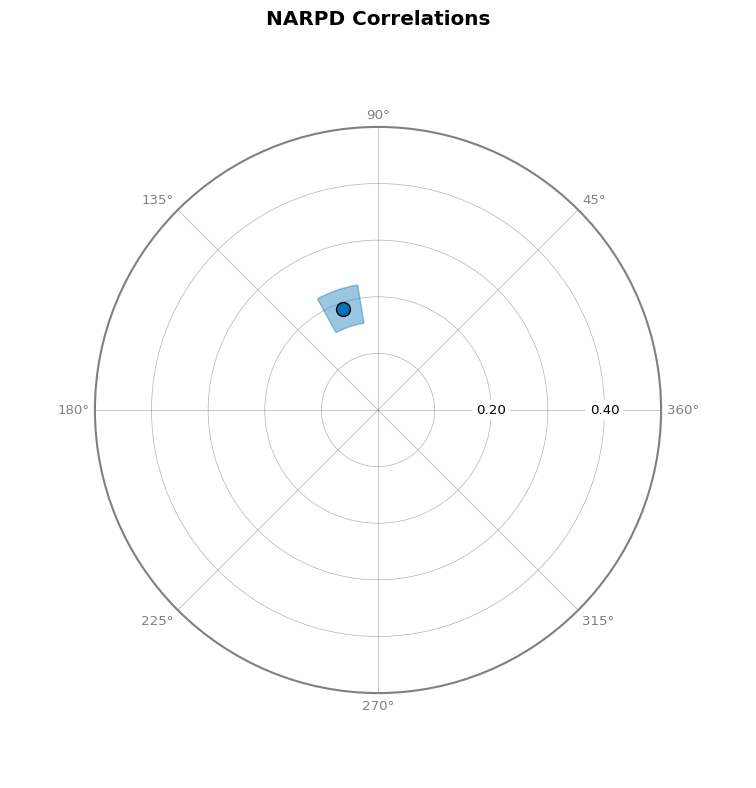
5. Group Comparisons¶
Multiple Groups¶
# Analyze by gender
results_group = ssm_analyze(
data=jz2017,
scales=scales,
angles=angles,
grouping="Gender",
)
results_group.summary()
Statistical Basis: Mean Scores Bootstrap Resamples: 2000 Confidence Level: 0.95 Listwise Deletion: True Scale Displacements: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0] Profile[Female] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.946 │ 0.908 │ 0.984 │ │ X-Value │ 0.459 │ 0.421 │ 0.497 │ │ Y-Value │ -0.31 │ -0.356 │ -0.268 │ │ Amplitude │ 0.554 │ 0.511 │ 0.599 │ │ Displacement │ 325.963 │ 321.939 │ 329.811 │ │ Model Fit │ 0.889 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[Male] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.884 │ 0.842 │ 0.926 │ │ X-Value │ 0.227 │ 0.192 │ 0.261 │ │ Y-Value │ -0.186 │ -0.225 │ -0.148 │ │ Amplitude │ 0.294 │ 0.258 │ 0.33 │ │ Displacement │ 320.685 │ 313.649 │ 328.014 │ │ Model Fit │ 0.824 │ │ │ └──────────────┴──────────┴──────────┴──────────┘
# Compare groups on circle plot
fig = results_group.plot_circle(colors="Set2", title="Interpersonal Problems by Gender")
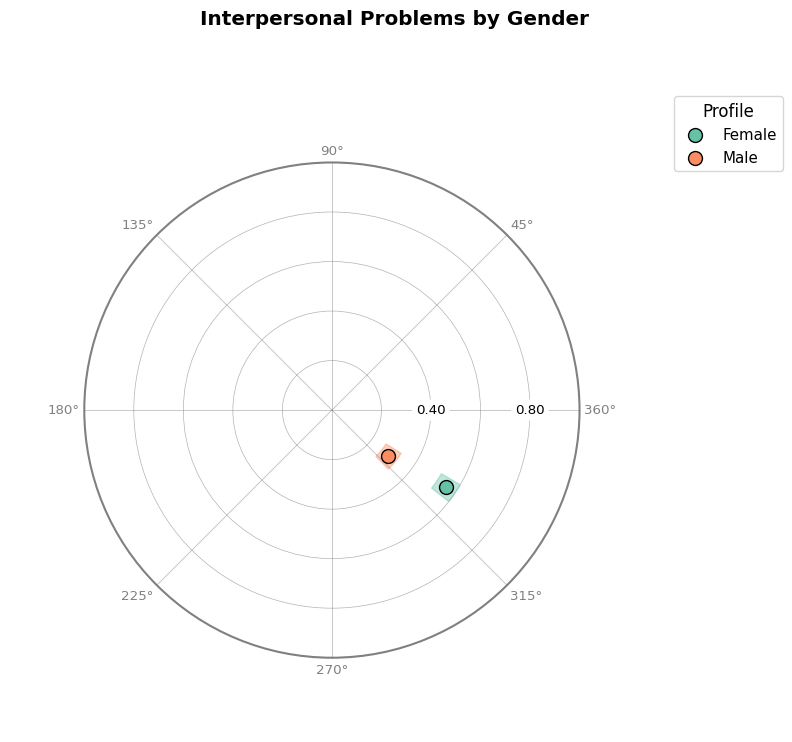
Multiple Measures¶
# Compare multiple personality disorder symptoms
results_multi = ssm_analyze(
data=jz2017,
scales=scales,
angles=angles,
measures=["NARPD", "ASPD", "BORPD"],
)
results_multi.summary()
Statistical Basis: Correlation Scores Bootstrap Resamples: 2000 Confidence Level: 0.95 Listwise Deletion: True Scale Displacements: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0] Profile[NARPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.202 │ 0.168 │ 0.238 │ │ X-Value │ -0.062 │ -0.094 │ -0.03 │ │ Y-Value │ 0.179 │ 0.143 │ 0.211 │ │ Amplitude │ 0.189 │ 0.154 │ 0.223 │ │ Displacement │ 108.967 │ 99.472 │ 118.732 │ │ Model Fit │ 0.957 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[ASPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.124 │ 0.089 │ 0.159 │ │ X-Value │ -0.099 │ -0.133 │ -0.062 │ │ Y-Value │ 0.203 │ 0.168 │ 0.237 │ │ Amplitude │ 0.226 │ 0.191 │ 0.262 │ │ Displacement │ 115.927 │ 106.85 │ 124.367 │ │ Model Fit │ 0.964 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[BORPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.277 │ 0.243 │ 0.31 │ │ X-Value │ -0.036 │ -0.069 │ -0.004 │ │ Y-Value │ 0.097 │ 0.056 │ 0.138 │ │ Amplitude │ 0.103 │ 0.064 │ 0.146 │ │ Displacement │ 110.358 │ 92.848 │ 130.012 │ │ Model Fit │ 0.872 │ │ │ └──────────────┴──────────┴──────────┴──────────┘
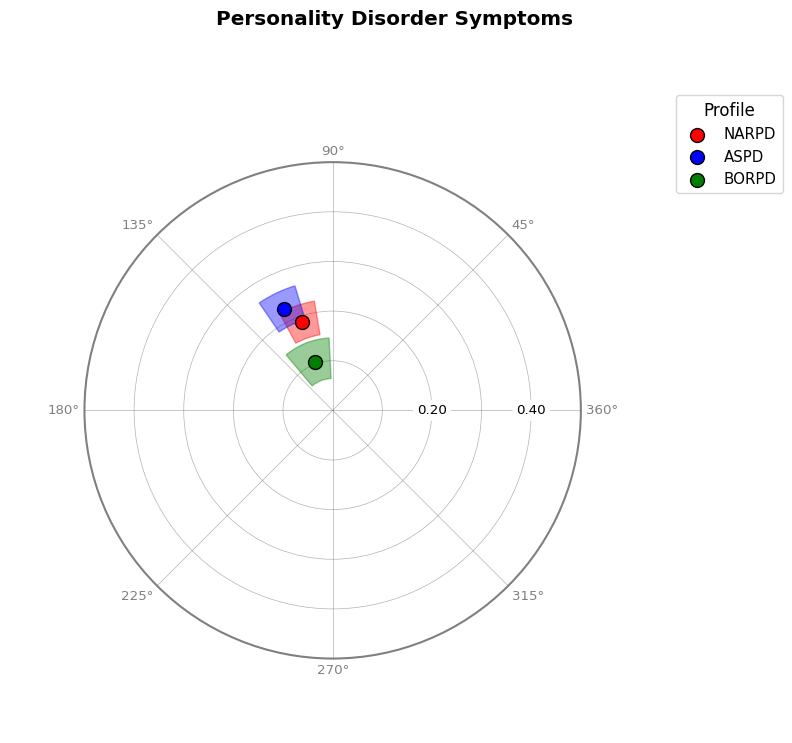
# Visualize multiple measures
fig = results_multi.plot_circle(
colors=["red", "blue", "green"], title="Personality Disorder Symptoms"
)
6. Contrast Analyses¶
Group Contrasts¶
To test whether groups differ significantly on SSM parameters, use
contrast=True:
# Contrast analysis between genders
results_contrast = ssm_analyze(
data=jz2017,
scales=scales,
angles=angles,
grouping="Gender",
contrast=True,
)
results_contrast.summary()
Statistical Basis: Mean Scores Bootstrap Resamples: 2000 Confidence Level: 0.95 Listwise Deletion: True Scale Displacements: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0] Profile[Female] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.946 │ 0.906 │ 0.983 │ │ X-Value │ 0.459 │ 0.422 │ 0.496 │ │ Y-Value │ -0.31 │ -0.352 │ -0.267 │ │ Amplitude │ 0.554 │ 0.512 │ 0.599 │ │ Displacement │ 325.963 │ 322.128 │ 329.684 │ │ Model Fit │ 0.889 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[Male] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.884 │ 0.842 │ 0.927 │ │ X-Value │ 0.227 │ 0.191 │ 0.263 │ │ Y-Value │ -0.186 │ -0.223 │ -0.147 │ │ Amplitude │ 0.294 │ 0.258 │ 0.33 │ │ Displacement │ 320.685 │ 313.831 │ 327.899 │ │ Model Fit │ 0.824 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[Male - Female] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ -0.062 │ -0.118 │ -0.004 │ │ X-Value │ -0.232 │ -0.282 │ -0.18 │ │ Y-Value │ 0.124 │ 0.068 │ 0.18 │ │ Amplitude │ -0.261 │ -0.317 │ -0.203 │ │ Displacement │ -5.278 │ 347.143 │ 2.851 │ │ Model Fit │ -0.066 │ │ │ └──────────────┴──────────┴──────────┴──────────┘
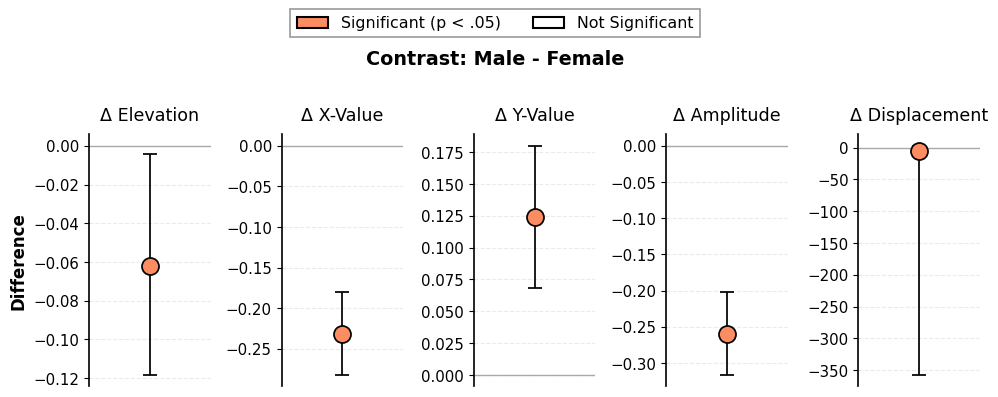
# View contrast results (last row shows Male - Female differences)
GT(results_contrast.results.round(2))
| Label | Group | Measure | e_est | e_lci | e_uci | x_est | x_lci | x_uci | y_est | y_lci | y_uci | a_est | a_lci | a_uci | d_est | d_lci | d_uci | fit_est |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Female | Female | 0.95 | 0.91 | 0.98 | 0.46 | 0.42 | 0.5 | -0.31 | -0.35 | -0.27 | 0.55 | 0.51 | 0.6 | 325.96 | 322.13 | 329.68 | 0.89 | |
| Male | Male | 0.88 | 0.84 | 0.93 | 0.23 | 0.19 | 0.26 | -0.19 | -0.22 | -0.15 | 0.29 | 0.26 | 0.33 | 320.68 | 313.83 | 327.9 | 0.82 | |
| Male - Female | Male - Female | -0.06 | -0.12 | -0.0 | -0.23 | -0.28 | -0.18 | 0.12 | 0.07 | 0.18 | -0.26 | -0.32 | -0.2 | -5.28 | 347.14 | 2.85 | -0.07 |
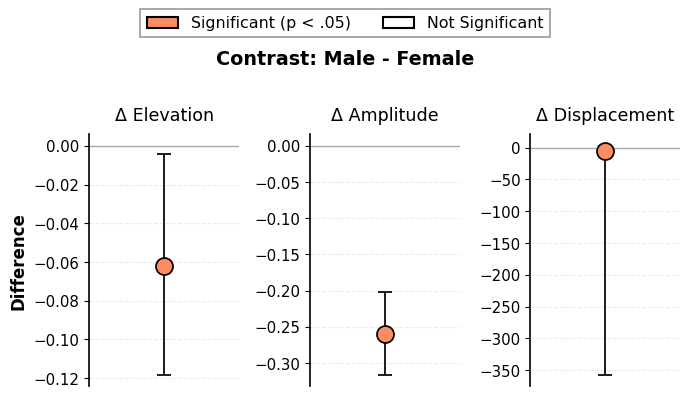
# Simplified contrast plot (drop X and Y parameters)
fig = results_contrast.plot_contrast(drop_xy=True)
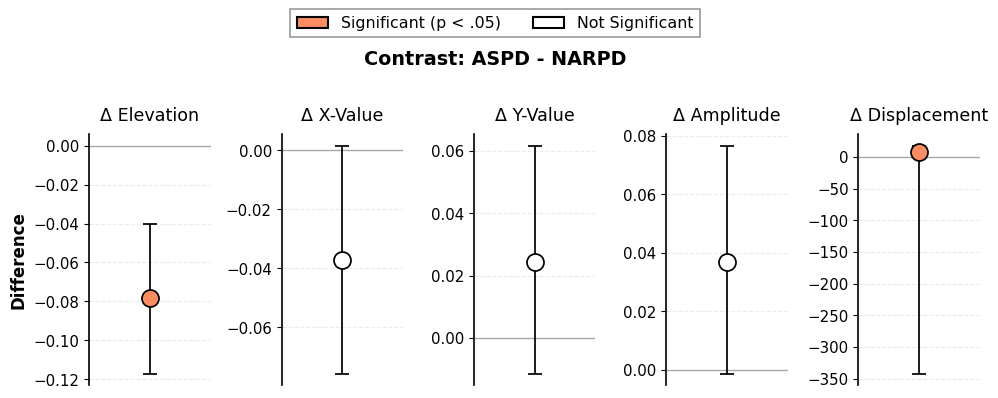
Measure Contrasts¶
# Compare two measures
results_meas_contrast = ssm_analyze(
data=jz2017,
scales=scales,
angles=angles,
measures=["NARPD", "ASPD"],
contrast=True,
)
results_meas_contrast.summary()
Statistical Basis: Correlation Scores Bootstrap Resamples: 2000 Confidence Level: 0.95 Listwise Deletion: True Scale Displacements: [90.0, 135.0, 180.0, 225.0, 270.0, 315.0, 360.0, 45.0] Profile[NARPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.202 │ 0.169 │ 0.235 │ │ X-Value │ -0.062 │ -0.094 │ -0.028 │ │ Y-Value │ 0.179 │ 0.146 │ 0.212 │ │ Amplitude │ 0.189 │ 0.156 │ 0.224 │ │ Displacement │ 108.967 │ 99.04 │ 118.521 │ │ Model Fit │ 0.957 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[ASPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ 0.124 │ 0.088 │ 0.16 │ │ X-Value │ -0.099 │ -0.135 │ -0.063 │ │ Y-Value │ 0.203 │ 0.169 │ 0.238 │ │ Amplitude │ 0.226 │ 0.19 │ 0.262 │ │ Displacement │ 115.927 │ 107.14 │ 124.123 │ │ Model Fit │ 0.964 │ │ │ └──────────────┴──────────┴──────────┴──────────┘ Profile[ASPD - NARPD] ┏━━━━━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┳━━━━━━━━━━┓ ┃ ┃ Estimate ┃ Lower CI ┃ Upper CI ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━╇━━━━━━━━━━┩ │ Elevation │ -0.079 │ -0.117 │ -0.04 │ │ X-Value │ -0.037 │ -0.076 │ 0.001 │ │ Y-Value │ 0.024 │ -0.012 │ 0.062 │ │ Amplitude │ 0.037 │ -0.001 │ 0.077 │ │ Displacement │ 6.96 │ 356.639 │ 17.454 │ │ Model Fit │ 0.007 │ │ │ └──────────────┴──────────┴──────────┴──────────┘
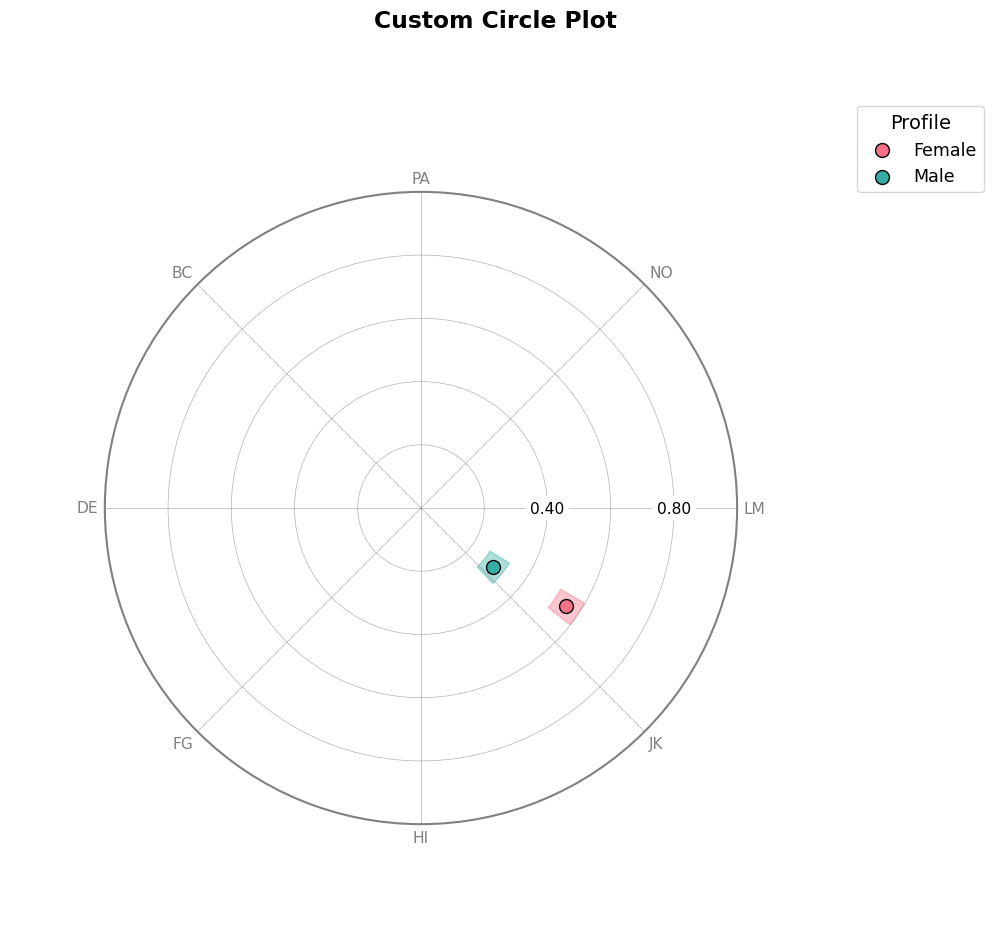
7. Customizing Visualizations¶
All plot functions support extensive customization:
# Circle plot with custom options
fig = results_group.plot_circle(
colors="husl", # Color palette
fontsize=14, # Font size
figsize=(10, 10), # Figure size
title="Custom Circle Plot",
angle_labels=scales, # Custom angle labels
)
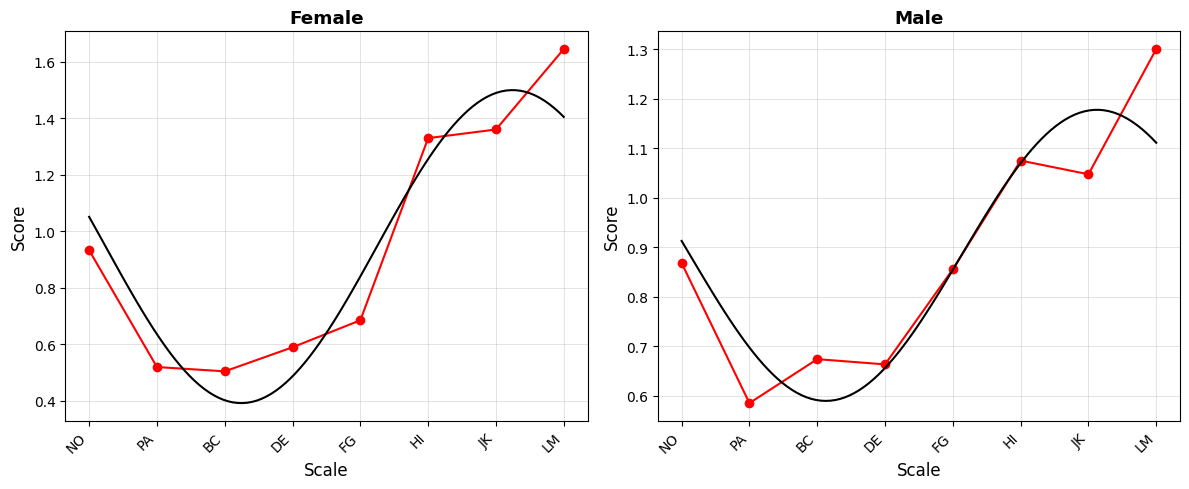
# Curve plot with custom options
fig = results_group.plot_curve(
angle_labels=scales,
base_size=12,
figsize=(12, 5),
)
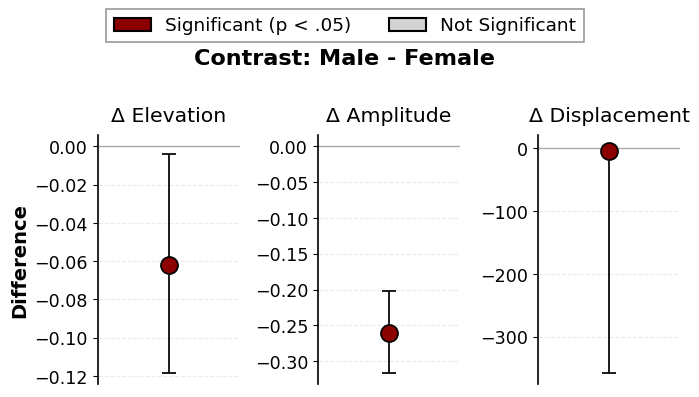
# Contrast plot with custom colors
fig = results_contrast.plot_contrast(
sig_color="darkred",
ns_color="lightgray",
fontsize=14,
drop_xy=True,
)
8. Saving Plots¶
All plots return matplotlib Figure objects that can be saved:
# Save plots
fig1 = results.plot_circle()
fig1.savefig("circle_plot.png", dpi=300, bbox_inches="tight")
fig2 = results.plot_curve()
fig2.savefig("curve_plot.png", dpi=300, bbox_inches="tight")
Summary¶
The circumplex package provides a comprehensive toolkit for SSM analysis:
ssm_analyze(): Performs mean-based or correlation-based SSM analysisplot_circle(): Visualizes amplitude/displacement on circular plotsplot_curve(): Shows fitted curves with observed dataplot_contrast(): Displays parameter differences between groups
All functions support: - Single or multiple groups - Single or multiple measures - Bootstrap confidence intervals - Contrast analyses - Extensive customization options